Tracking Experiments and Visualizing Results
While an experiment is running, and any time after it finishes, track it and visualize the results in the ClearML Web UI, including:
- Execution details - Code, the base Docker image used for ClearML Agent, output destination for artifacts, and the logging level.
- Configuration - Hyperparameters, user properties, and configuration objects.
- Artifacts - Input model, output model, model snapshot locations, other artifacts.
- General information - Information about the experiment, for example: the experiment start, create, and last update times and dates, user creating the experiment, and its description.
- Console - stdout, stderr, output to the console from libraries, and ClearML explicit reporting.
- Scalars - Metric plots.
- Plots - Other plots and data, for example: Matplotlib, Plotly, and ClearML explicit reporting.
- Debug samples - Images, audio, video, and HTML.
Viewing Modes
The ClearML Web UI provides two viewing modes for experiment details:
The info panel
Full screen details mode.
Both modes contain all experiment details. When either view is open, switch to the other mode by clicking
(View in experiments table / full screen), or clicking
(menu) > View in experiments
table / full screen.
Info Panel
The info panel keeps the experiment table in view so that experiment actions can be performed from the table (as well as the menu in the info panel).
![]()
Click to
hide details in the experiment table, so only the experiment names and statuses are displayed
![]()
Full Screen Details View
The full screen details view allows for easier viewing and working with experiment tracking and results. The experiments table is not visible when the full screen details view is open. Perform experiment actions from the menu.
![]()
Execution
An experiment's EXECUTION tab of lists the following:
- Source code
- Uncommitted changes
- Installed Python packages
- Container details
- Output details
In full-screen mode, the source code and output details are grouped in the DETAILS section.
Source Code
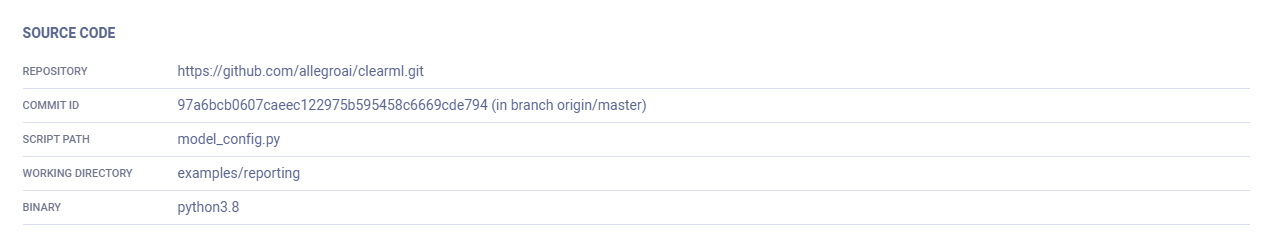
The Source Code section of an experiment's EXECUTION tab includes:
- The experiment's repository
- Commit ID
- Script path
- Working directory
- Binary (Python executable)
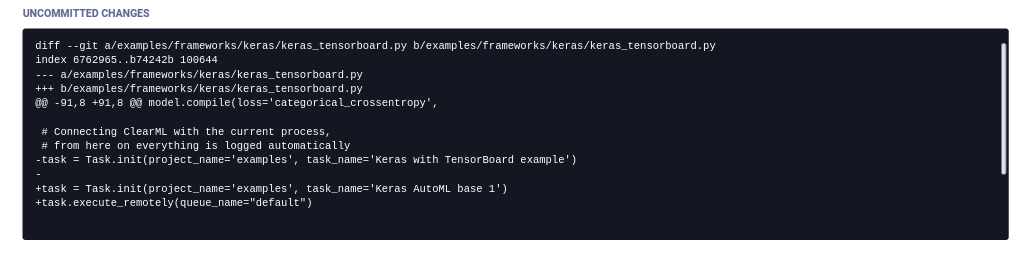
Uncommitted Changes
ClearML displays the git diff of the experiment in the Uncommitted Changes section.
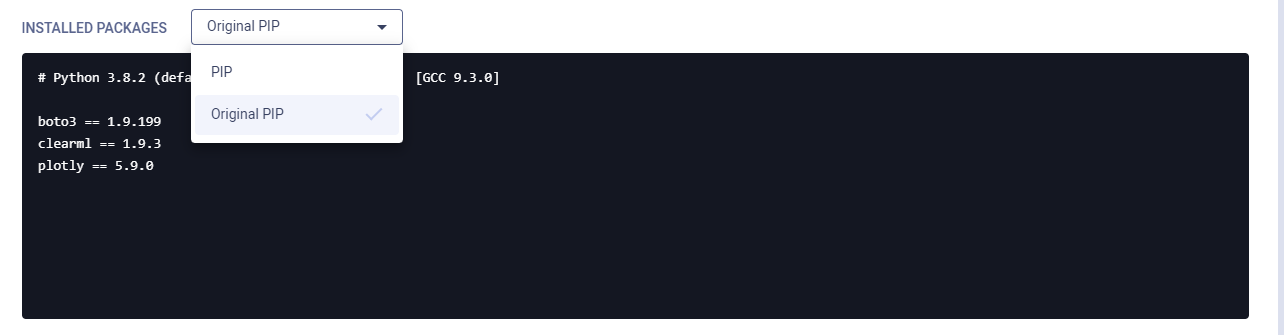
Installed Packages
The Installed Packages section lists the experiment's installed Python packages and their versions.

When a ClearML agent executing an experiment ends up using a different set of python packages than was originally
specified, both the original specification (original pip or original conda), and the packages the agent ended up
using to set up an environment (pip or conda) are available. Select which requirements to view in the dropdown menu.
Container
The Container section list the following information:
- Image - a pre-configured Docker that ClearML Agent will use to remotely execute this experiment (see Building Docker containers)
- Arguments - add Docker arguments
- Setup shell script - a bash script to be executed inside the Docker before setting up the experiment's environment

Output
The Output details include:
- The output destination used for storing model checkpoints (snapshots) and artifacts (see also, default_output_uri
in the configuration file, and
output_uriinTask.initparameters).
Configuration
All parameters and configuration objects appear in the CONFIGURATION tab.
Hyperparameters
Hyperparameters are grouped by their type and appear in CONFIGURATION > HYPERPARAMETERS. Once an experiment is run and stored in ClearML Server, any of these hyperparameters can be modified.
Command Line Arguments
The Args group shows automatically logged argument parser parameters (e.g. argparse, click, hydra).
Hover over (menu) on a
parameter's line, and the type, description, and default value appear, if they were provided.
![]()
Environment Variables
If the CLEARML_LOG_ENVIRONMENT variable was set, the Environment group will show environment variables (see this FAQ).
![]()
Custom Parameter Groups
Custom parameter groups show parameter dictionaries if the parameters were connected to the Task, using
Task.connect() with a name argument provided. General is the default section
if a name is not provided.
![]()
TensorFlow Definitions
The TF_DEFINE parameter group shows automatic TensorFlow logging.
![]()
User Properties
User properties allow to store any descriptive information in a key-value pair format. They are editable in any experiment, except experiments whose status is Published (read-only).
![]()
Configuration Objects
ClearML tracks experiment (Task) model configuration objects, which appear in Configuration Objects > General.
These objects include those that are automatically tracked, and those connected to a Task in code (see Task.connect_configuration).
![]()
ClearML supports providing a name for a Task model configuration object (see the name
parameter in Task.connect_configuration).
![]()
Artifacts
Artifacts tracked in an experiment appear in the ARTIFACTS tab, and include models and other artifacts.
Artifacts location is stored in the FILE PATH field.
The UI provides locally stored artifacts with a 'copy to clipboard' action ()
to facilitate local storage access (since web applications are prohibited from accessing the local disk for security reasons).
The UI provides Network hosted (e.g. https://, s3:// etc. URIs) artifacts with a download action (
)
to retrieve these files.
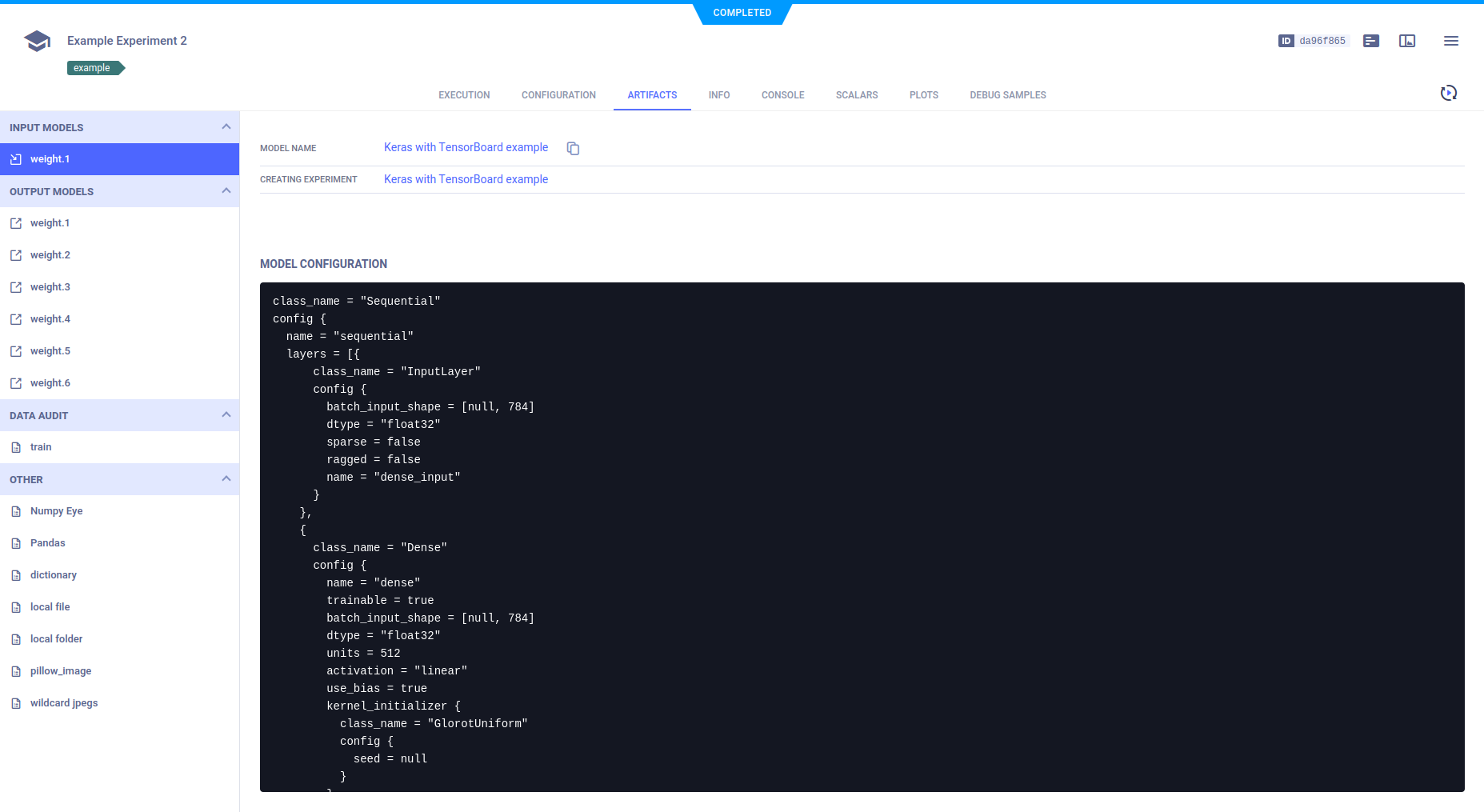
Models
The input and output models appear in the ARTIFACTS tab. Models are associated with the experiment, but to see further model details, including design, label enumeration, and general information, go to the MODELS tab, by clicking the model name, which is a hyperlink to those details.
To retrieve a model:
In the ARTIFACTS tab > MODELS > Input Model or Output Model, click the model name hyperlink.
In the model details > GENERAL tab > MODEL URL, either:
- Download the model
, if it is stored in remote storage.
- Copy its location to the clipboard
, if it is in a local file.
- Download the model
Other Artifacts
Other artifacts, which are uploaded but not dynamically tracked after the upload, appear in the OTHER section. They include the file path, file size, and hash.
To retrieve Other artifacts:
In the ARTIFACTS tab > OTHER > Select an artifact > Either:
- Download the artifact
, if it is stored in remote storage.
- Copy its location to the clipboard
, if it is in a local file.
![]()
General Information
General experiment details appear in the INFO tab. This includes information describing the stored experiment:
- The parent experiment
- Project name
- Creation, start, and last update dates and times
- User who created the experiment
- Experiment state (status)
- Whether the experiment is archived
- Runtime properties - Information about the machine running the experiment, including:
- Operating system
- CUDA driver version
- Number of CPU cores
- Number of GPUs
- CPU / GPU type
- Memory size
- Host name
- Processor
- Python version
- Experiment Progress
![]()
Experiment Results
You can embed experiment plots and debug samples into ClearML Reports. These visualizations are updated live as the experiment(s) updates. The Enterprise Plan and Hosted Service support embedding resources in external tools (e.g. Notion). See Plot Controls.
Console
The complete experiment log containing everything printed to stdout and stderr appears in the CONSOLE tab. The full log is downloadable. To view the end of the log, click Jump to end.
![]()
Scalars
All scalars that ClearML automatically logs, as well as those explicitly reported in code, appear in SCALARS.
Scalar series are presented in a line chart. To see the series for a metric in high resolution,
view it in full screen mode by hovering over the graph and clicking .
Scalar graphs in full screen mode do not auto-refresh. Click
to update the graph.
Reported single value scalars are aggregated into a table plot displaying scalar names and values (see Logger.report_single_value).
Scalar Plot Tools
Use the scalar tools to improve analysis of scalar metrics. In the info panel, click to use the tools. In the full screen details view, the tools
are on the left side of the window. The tools include:
Group by - Select one of the following:
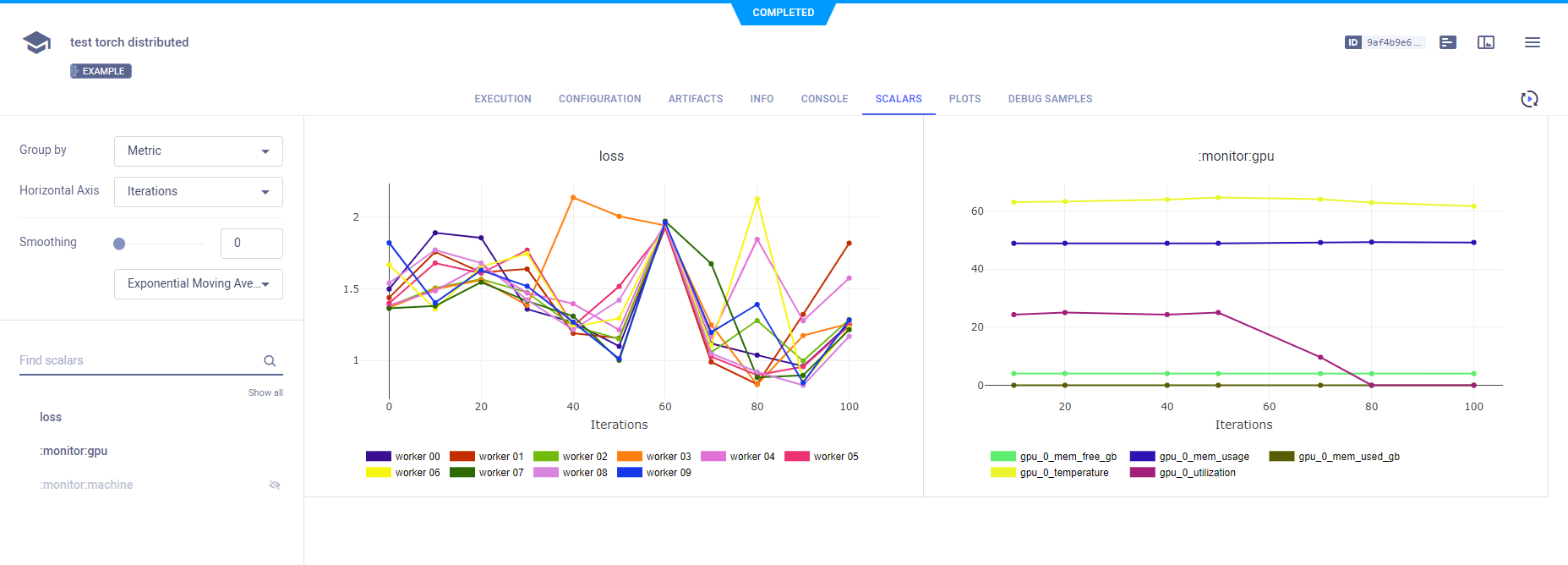
Metric - All variants for a metric on the same plot
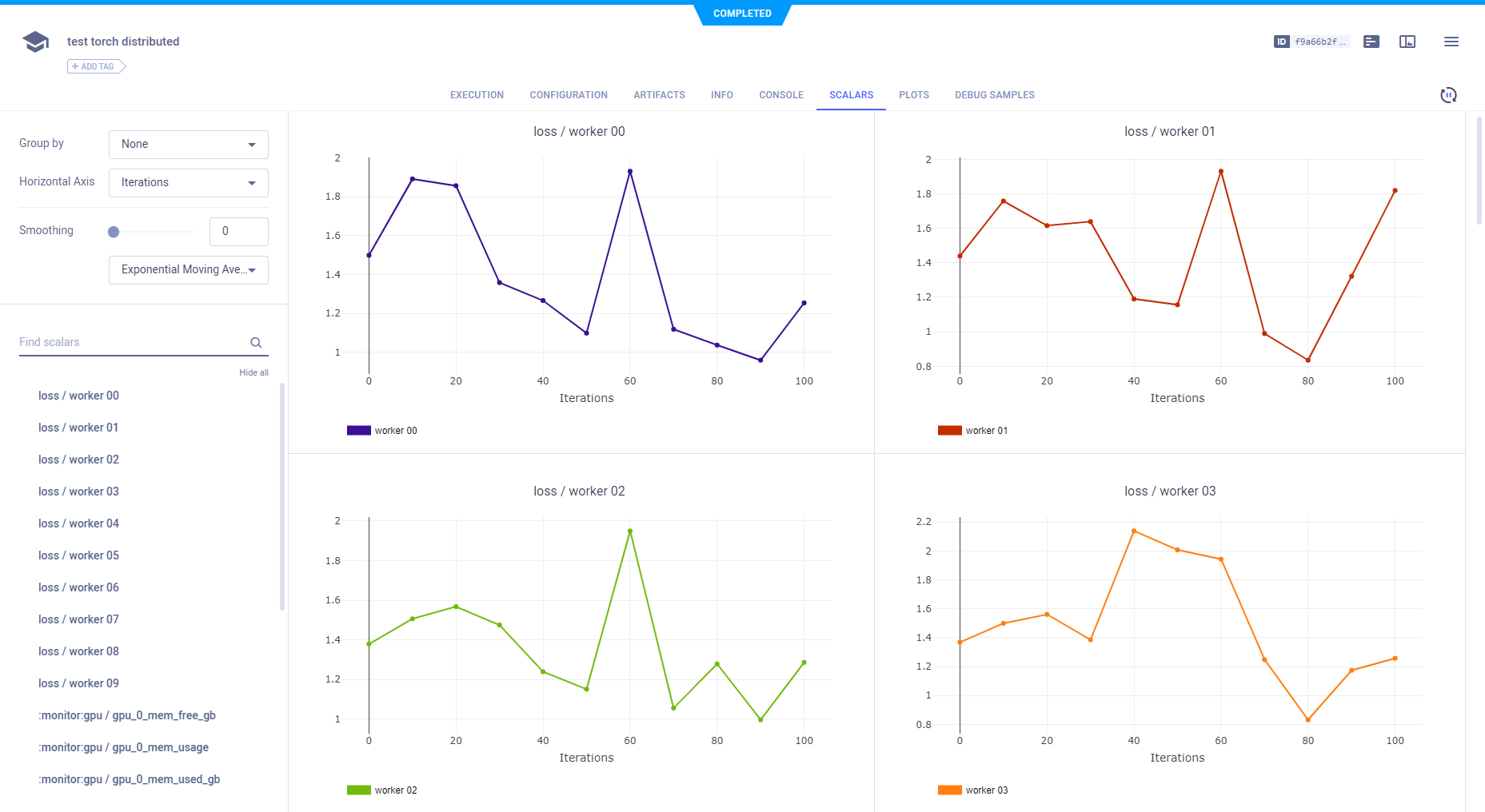
None - Group by metric and variant (individual metric-variant plots).
Horizontal axis - Select the x-axis units:
- Iterations
- Time from start - Time since experiment began
- Wall time - Local clock time
Curve smoothing - Choose which smoothing algorithm to use from the dropdown menu: Exponential moving average, Gaussian, or Running Average. Use the slider to configure the smoothing factor or specify a value manually.
Show / hide plots - Click HIDE ALL, and then click
on those you want to see.
To embed scalar plots in your Reports, hover over a plot and click ,
which will copy to clipboard the embed code to put in your Reports. In contrast to static screenshots, embedded resources
are retrieved when the report is displayed allowing your reports to show the latest up-to-date data.
See additional plot controls below.
Plots
Non-time-series plots appear in PLOTS. These include data generated by libraries, visualization tools, and explicitly reported using the ClearML Logger. These may include 2D and 3D plots, tables (Pandas and CSV files), and Plotly plots. Individual plots can be shown / hidden or filtered by title.
![]()
For each metric, the latest reported plot is displayed.
When viewing a plot in full screen (),
older iterations are available through the iteration slider (or using the up/down arrow keyboard shortcut). Go to the
previous/next plot in the current iteration using the
/
buttons (or using the left/right arrow keyboard shortcut).
![]()
Plot Controls
The table below lists the plot controls which may be available for any plot (in the SCALARS and PLOTS tabs). These controls allow you to better analyze the results. Hover over a plot, and the controls appear.
| Icon | Description |
|---|---|
| Download plots as PNG files. | |
| Pan around plot. Click | |
| To examine an area, draw a dotted box around it. Click | |
| To examine an area, draw a dotted lasso around it. Click | |
| Zoom into a section of a plot. Zoom in - Click | |
| Zoom in. | |
| Zoom out. | |
| Reset to autoscale after zooming ( | |
| Reset axes after a zoom. | |
| Show / hide spike lines. | |
Set data hover mode:
| |
| Switch to logarithmic view. | |
| Hide / show the legend. | |
| Switch between original and auto-fitted plot dimensions. The original layout is the plot's user-defined dimensions. | |
| Download plot data as a JSON file. | |
| Download table plot data as a CSV file. | |
| Expand plot to entire window. When used with scalar graphs, full screen mode displays plots with all data points, as opposed to an averaged plot | |
| Refresh scalar graphs in full screen mode to update it. | |
Copy to clipboard the resource embed code. This opens the following options:
|
3D Plot Controls
| Icon | Description |
|---|---|
| Switch to orbital rotation mode - rotate the plot around its middle point. | |
| Switch to turntable rotation mode - rotate the plot around its middle point while constraining one axis | |
| Reset axes to default position. |
Debug Samples
Experiment outputs such as images, audio, and videos appear in DEBUG SAMPLES. These include data generated by libraries and visualization tools, and explicitly reported using the ClearML Logger.
You can view debug samples by metric in the reported iterations. Filter the samples by metric by selecting a metric from the dropdown menu above the samples. The most recent iteration appears first.
![]()
For each metric, the latest reported debug sample is displayed.
Click a sample to view it in full screen. If the sample is video or audio, the full screen mode includes a player.
When viewing a sample in full screen, older iterations are available through the iteration slider (or using the up/down
arrow keyboard shortcut). Go to the previous/next sample in the current iteration using the /
buttons (or using the left/right arrow keyboard shortcut).
![]()
Tagging Experiments
Tags are user-defined, color-coded labels that can be added to experiments (and pipelines, datasets, and models), allowing to easily identify and group experiments. Tags can help in organizing, querying, and automating experiments. For example, tag experiments by the machine type used to execute them, label versions, team names, or any other category.
You can use tags to filter your experiments in your experiment table (see Filtering Columns) or when querying experiments in your code (see Tag Filters). You can trigger experiment execution according to their tags (see TriggerScheduler) or automatically deploy models according to their tags (see ClearML Serving).
To add tags:
- Click the experiment > Hover over the tag area > +ADD TAG or
(menu)
- Do one of the following:
- Add a new tag - Type the new tag name > (Create New).
- Add an existing tag - Click a tag.
- Customize a tag's colors - Click Tag Colors > Click the tag icon > Background or Foreground > Pick a color > OK > CLOSE.
To remove a tag - Hover over the tag and click X.
Locating the Experiment (Task) ID
The task ID appears in the experiment page's header.